Protein Structural Analysis Laboratory Publications
Michigan State University

Protein Structural Analysis Laboratory
Publications
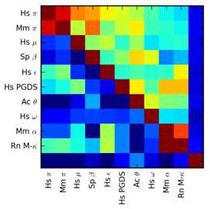 Joseph Bemister-Buffington, Alex J. Wolf, Sebastian Raschka and Leslie A. Kuhn (2020) "Machine Learning to Identify Flexibility Signatures of Class A GPCR Inhibition", Biomolecules, in press.
Joseph Bemister-Buffington, Alex J. Wolf, Sebastian Raschka and Leslie A. Kuhn (2020) "Machine Learning to Identify Flexibility Signatures of Class A GPCR Inhibition", Biomolecules, in press.J. Chen and L. A. Kuhn (2019) "Deciphering the Three-Domain Architecture in Schlafens and the Structures and Roles of Human Schlafen12 and SerpinB12 in Transcriptional Regulation", J. Molec. Graphics Modelling 90, 59-76. https://doi.org/10.1016/j.jmgm.2019.04.003
Supplementary Data 1 (hSLFN12 N-terminal domain PDB)
Supplementary Data 2 (hSLFN12 C-terminal domain PDB)
Supplementary Data 3 (hSerpinB12 PDB)
Supplementary Data 4 (Additional GTPase-like domain functions) S. Raschka, A. J. Wolf, J. Bemister-Buffington, and L. A. Kuhn (2018) "Protein-ligand interfaces are polarized: Discovery of a strong trend for intermolecular hydrogen bonds to favor donors on the protein side with implications for predicting and designing ligand complexes", J. Comp.-Aided Molec. Design 32(4), 511-528. S. Raschka, A. M. Scott, N. Liu, S. Gunturu, M. Huertas, W. Li, and L. A. Kuhn (2018) "Enabling the hypothesis-driven prioritization of ligand candidates in big databases: Screenlamp and its application to GPCR inhibitor discovery for invasive species control", J. Comp.-Aided Molec. Design 32(3), 415-433. Supplementary Materials S. Raschka, A. M. Scott, M. Huertas, W. Li, and L. A. Kuhn (2018) "Automated Inference of Chemical Discriminants of Biological Activity", Methods in Molecular Biology: Computational Drug Discovery and Design (M. Gore and U.B. Jagtap, ed.), Springer Protocols, Humana Press, New York, NY, pp. 307-338. S. Raschka, S. K. More, D. Devadoss, B. Zeng, L. A. Kuhn*, M. D. Basson* (2018) "Identification of potential small-molecule protein-protein inhibitors of cancer metastasis by 3D epitope-based computational screening", Journal of Physiology and Pharmacology, in press. *Joint corresponding authors M. D. Basson, Q. Wang, L. S. Chaturvedi, S. More, E. E. Vomhof-DeKrey, S. Al-Marsoummi, K. Sun, L. A. Kuhn, P. Kovalenko, and M. Kiupel (2018) "Schlafen 12 interaction with Serpin B12 and deubiquitylases drives human enterocyte differentiation", Cellular Physiology and Biochemistry, 48(3):1274-1290. B. Zeng, D. Devadoss, S. Wang, E. DeKrey, L. A. Kuhn, and M. D. Basson (2017) "Inhibition of pressure-activated cancer cell adhesion by FAK-derived peptides", Oncotarget 8, 98051-98067. D. D. McKenna, E. D. Scully, Y. Pauchet, K. Hoover, R. Kirsch, S. M. Geib, R. F. Mitchell, R. M. Waterhouse, S.-J. Ahn, D. Arsala, A. K. Bennett, J. B. Benoit, H. Blackmon, T. Bledsoe, J. H. Bowsher, A. Busch, B. Calla, H. Chao, C. Childers, D. J. Clarke, Lorna Cohen, Jeff Demuth, Huyen Dinh, HarshaVardhan Doddapaneni, Amanda Dolan, J. J. Duan, S. Dugan, M. Friedrich, K. M. Glastad, M.A.D. Goodisman, S. Haddad, Y. Han, D. S. T. Hughes, P. Ioannidis, I. M. Vargas Jentzsch, J. S. Johnston, J. W. Jones, L. A. Kuhn, D. R. Lance, C.-Y. Lee, S. L. Lee, H. Lin, J. A. Lynch, A. P. Moczek, S. C. Murali, D. M. Muzny, D. R. Nelson, S. R. Palli, K. A. Panfilio, D. Pers, M. Poelchau, H. Quan, J. Qu, A. M. Ray, J. P. Rinehart, H. M. Robertson, R. Roehrdanz, A. J. Rosendale, S. Shin, C. Silva, A. C. Torson, J. H. Werren, K. C. Worley, G. Yocum, E. M. Zdobnov, R. A. Gibbs, and S. Richards (2016) "The Asian longhorned beetle (Anoplophora glabripennis) genome reveals key functional and evolutionary innovations at the beetle-plant interface", Genome Biology 17, 227. Supplementary Information S. Raschka, J. Bemister-Buffington, and L. A. Kuhn (2016) "Detecting the Native Ligand Orientation by Interfacial Rigidity: SiteInterlock (with Supporting Information)", Proteins: Structure, Function, and Bioinformatics, 84(12) 1888-1901. F. Li, J. Liu, N. Liu, C. Hiser, L.A. Kuhn, R.M. Garavito and S. Ferguson-Miller (2016) "Translocator protein 18 kDa (TSPO): An old protein with new functions?", Biochemistry, 55 (20), 2821-2831. N. Liu, J. R. Van Voorst, J. Johnston, and L. A. Kuhn (2015) "CholMine: Determinants and prediction of cholesterol and cholate binding across non-homologous protein structures", J. Chemical Information and Modeling, 55(4), 747-759, DOI:10.1021/ci5006542. Supporting Information (pdf) C. P. Najt, J. S. Lwande, A. L. McIntosh, S. Senthivinayagam, S. Gupta, L. A. Kuhn, and B. P. Atshaves (2014) "Structural and Functional Assessment of Perilipin 2 Lipid Binding Domain(s)" (pdf)
Supporting Information (pdf) R. L. Roston, K. Wang, L. A. Kuhn, and C. Benning (2014) "Structural determinants allowing transferase activity in SENSITIVE TO FREEZING 2, classified as a family I glycosyl hydrolase" (pdf)
Supplementary Materials(pdf) C. Hiser, L. Buhrow, J. Liu, L. Kuhn and S. Ferguson-Miller (2013) "A Conserved Amphipathic Ligand Binding Region Influences K-Path Dependent Activity of Cytochrome c Oxidase" (pdf) Biochemistry 52(8), 1385-1396. J. Van Voorst, Y. Tong, and L. A. Kuhn (2012) "ArtSurf: A Method for Deformable Partial Matching of Protein Small-Molecule Binding Sites" (pdf) Proceedings of the ACM Conference on Bioinformatics, Computational Biology and Biomedicine, 36-43. M. A. Kron, C. Wang, S. Vodanovich-Jankovic, O. M. Z. Howard, and L. A. Kuhn (2012) "Interleukin-8-like activity in a filarial asparaginyl-tRNA synthetase" (pdf) Molecular & Biochemical Parasitology, 185, 66-69.
Supplementary Materials (doc) L. Buhrow, S. Ferguson-Miller and L. A. Kuhn (2012) "From Static Structure to Living Protein: Computational Analysis of Cytochrome c Oxidase Main-chain Flexibility" (pdf) Biophysical Journal, 102, 2158-2166.
Supporting Materials (pdf) D. Canella, S. J. Gilmour, L. A. Kuhn, and M. F. Thomashow (2010) "DNA binding by the Arabidopsis CBF1 transcription factor requires the PKKP/RAGRxKFxETRHP signature sequence." (pdf) Biochim Biophys Acta, 1799(5-6), 454-62.
M. I. Zavodszky, A. Rohatgi, J. R. Van Voorst, H. Yan, and L. A. Kuhn (2009) "Scoring Ligand Similarity in Structure-Based Virtual Screening" (pdf) Journal of Molecular Recognition, 22(4), 280-292.
(Figures) (pdf)
Supplementary Materials (pdf) K. S. Keating, S. C. Flores, M. B. Gerstein, and L. A. Kuhn (2009) "StoneHinge: Hinge Prediction by Network Analysis of Individual Protein Structures" (pdf) Protein Science 18, 359-371. S. Quiroz, S. C. K. Sukuru, R. P. Hausinger, L. A. Kuhn, and W. T. Heller (2008) "The Structure of Urease Activation Complexes Examined by Flexibility Analysis, Mutagenesis, and Small-Angle X-Ray Scattering Approaches" (pdf) Arch. Biochem. Biophys. 480, 51-57. P. Cozzini, G. E. Kellogg, F. Spyrakis, D. J. Abraham, G. Costantino, A. Emerson, F. Fanelli, H. Gohlke, L. A. Kuhn, G. M. Morris, M. Orozco, T. A. Pertinhez, M. Rizzi, and C. A. Sotriffer (2008) "Target Flexibility: An Emerging Consideration in Drug Discovery and Design" (pdf) J. Med. Chem. 51, 6237-6255. S. C. Flores, K. S. Keating, J. Painter, F. G. Morcos, K. Nguyen, E. A. Merritt, L. A. Kuhn, and M. B. Gerstein (2008) "HingeMaster: Normal Mode Hinge Prediction Approach and Integration of Complementary Predictors" (pdf) Proteins 73, 299-319. M. A. Kron, S. Kutlesa, A. Hendrick, A. Liu, J. Leykam, S. Cichenowicz, and L. A. Kuhn (2008) "Using Structural Analysis to Generate Parasite-Selective Monoclonal Antibodies" (pdf) Protein Science 17, 983-989. L. A. Kuhn (2008) "Strength in Flexibility: Modeling Side-chain Conformational Change in Docking and Screening" (pdf) Structure-Based Drug Discovery (R. M. Stroud and J. Finer-Moore, Eds.), Royal Society of Chemistry, London, pp. 177-187. T. A. Muller, M. I. Zavodszky, M. Feig, L. A. Kuhn, and R. P. Hausinger (2006) "Structural Basis for the Enantiomeric Specificities of R- and S-specific Phenoxypropionate/Alpha-ketoglutarate Dioxygenases" (pdf) Protein Science 15, 1356-1368.
(Figures) (pdf)
Supplementary Materials (pdf) S. C. K. Sukuru, T. Crepin, Y. Milev, L. C. Marsh, J. B. Hill, R. J. Anderson, J. C. Morris, A. Rohatgi, G. O'Mahony, M. Grotli, F. Danel, Y. Milev, M. G. P. Page, M. Hartlein, S. Cusack, M. A. Kron, and L. A. Kuhn (2006) "Discovering New Classes of Brugia malayi Asparaginyl tRNA Synthetase Inhibitors and Relating Specificity to Conformational Change" (pdf) J. Comp.-Aided Molec. Design 20, 159-178.
Supplementary Materials (pdf) M.I. Zavodszky, L.A. Kuhn (2005) "Side-Chain Flexibility in Protein-Ligand Binding: The Minimal Rotation Hypothesis" (pdf) Protein Science, 14, 1104 - 1114. P.-P. Kung, L. R. Zehnder, J. J. Meng, S. W. Kupchinsky, D. J. Skalitzky, M. C. Johnson, K. A. Maegley, A. Ekker, L. A. Kuhn, P. W. Rose, and L. A. Bloom (2005) "Design, Synthesis, and Biological Evaluation of Novel Human 5'-Deoxy-5'-Methylthioadenosine Phosphorylase (MTAP) Substrates" (pdf) Bioorg. Med. Chem. Letters 15, 2829-2833. M. Lei, M.I. Zavodszky, L.A. Kuhn, M.F. Thorpe (2004) "Sampling Protein Conformations and Pathways" (pdf) J Comp Chem, 25(9):1133-48. M. I. Zavodszky, M. Lei, A. R. Day, M. F. Thorpe, and L. A. Kuhn (2004) "Modeling Correlated Main-chain Motions in Proteins for Flexible Molecular Recognition" (pdf) Proteins: Struct. Funct. Bioinf., 57(2):243-261. H. Gohlke, L. A. Kuhn, and D. A. Case (2004) "Change in Protein Flexibility Upon Complex Formation: Analysis of Ras-Raf Using Molecular Dynamics and Molecular Approach" (pdf) Proteins: Struct. Funct. Bioinf., 56:322-337. M. A. Kron, L. A. Kuhn, P. C. Sanschagrin, M. Hartlein, and S. Cusak (2003) "Strategies for Antifilarial Drug Development" (pdf) J. Parasitology, 89, S226-S235. M. L. Raymer, L. A. Kuhn, and W. F. Punch (2003) "Knowledge Discovery in Biological Datasets Using a Hybrid Bayes Classifier/Evolutionary Algorithm" (pdf) IEEE Symposium on Systems, Man, and Cybernetics, 1-12. M. I. Zavodszky, P. C. Sanschagrin, R. S. Korde, and L. A. Kuhn (2002) "Distilling the Essential Features of a Protein Surface for Improving Protein-Ligand Docking, Scoring, and Virtual Screening" (pdf) J. Comp.-Aided Molecular Design, 16, 883-902. B. M. Hespenheide and L. A. Kuhn (2003) "Discovery of a Significant, Non-topological Preference for Antiparallel Alignment of Helices with Parallel Regions in Sheets" (pdf) Protein Science, 12, 1119-1125. B. M. Hespenheide, A. J. Rader, M. F. Thorpe, and L. A. Kuhn (2002) "Identifying Protein Folding Cores: Observing the Evolution of Rigid and Flexible Regions During Unfolding" (pdf)J. Molec. Graphics and Modelling, 21, 195-207. A. J. Rader, B. M. Hespenheide, L. A. Kuhn, and M. F. Thorpe (2002) "Protein Unfolding: Rigidity Lost" (pdf) Proceedings of the National Academy of Sciences USA 99, 3540-3545. E. E. Howell, U. Shukla, S. N. Hicks, R. D. Smiley, L. A. Kuhn, and M. I. Zavodszky (2001) "One Site Fits Both: A Model for the Ternary Complex of Folate + NADPH in R67 Dihydrofolate Reductase, a D2 Symmetric Enzyme" (pdf) J. Computer-Aided Molec. Design, 15, 1035-1052. D. J. Jacobs, A. J. Rader, L. A. Kuhn, and M. F. Thorpe (2001) "Protein Flexibility Predictions Using Graph Theory" (pdf) Proteins: Structure, Function, and Genetics 44, 150-165. M. F. Thorpe, M. Lei, A. J. Rader, D. J. Jacobs, and L. A. Kuhn (2001) "Protein Flexibility and Dynamics Using Constraint Theory" (pdf) J. Molec. Graphics and Modelling, 19, 60-69. M. F. Thorpe, M. Lei, A. J. Rader, D. J. Jacobs, and L. A. Kuhn (2001) "Protein Flexibility and Dynamics Using Constraint Theory" (pdf) in Biological Modeling: Protein Flexibility and Folding (L. A. Kuhn and M. F. Thorpe, eds.), Elsevier, New York, pp. 60-69. V. Schnecke and L. A. Kuhn (2000) "Virtual Screening with Solvation and Ligand-Induced Complementarity" (pdf) Perspectives in Drug Design and Discovery, 20, 171-190. M. L. Raymer, W. F. Punch, E. D. Goodman, L. A. Kuhn, and A. K. Jain (2000) "Dimensionality Reduction Using Genetic Algorithms" (pdf) IEEE Transactions on Evolutionary Computation, 4, 164-171. M. F. Thorpe, B. M. Hespenheide, Y. Yang, and L. A. Kuhn (2000) "Flexibility and Critical Hydrogen Bonds in Cytochrome c" (pdf) Pacific Symposium on Biocomputing, eds. R. B. Altman, A. K. Dunker, L. Hunter, K. Lauderdale, & T. Klein, World Scientific, Singapore, pp. 191-205. M. F. Thorpe and L. A. Kuhn (1999) "Rigidity in Glasses and Proteins" (pdf) Periodica Mathematica Hungarica 39, 241-252. L. Fan, P. C. Sanschagrin, L. S. Kaguni, and L. A. Kuhn (1999) "The Accessory Subunit of Mitochondrial DNA Polymerase Shares Structural Homology with Aminoacyl-tRNA Synthetases: Implications for a Dual Role as a Primer Recognition Factor and Processivity Clamp" (pdf) Proceedings of the National Academy of Sciences, USA 96, 9527-9532. B. Essigmann, B. Hespenheide, L. A. Kuhn, and C. Benning (1999) "Prediction of the Active-Site Structure and NAD+ Binding in SQD1, a Protein Essential for Sulfolipid Biosynthess in Arabidopsis" (pdf) Arch. Biochem. Biophys. 369, 30-41. V. Schnecke and L. A. Kuhn (1999) "Database Screening for HIV Protease Ligands: The Influence of Binding-Site Conformation and Representation on Ligand Selectivity" (pdf) Proceedings of the Seventh International Conference in Intelligent Systems for Molecular Biology, AAAI Press, pp. 242-251. Q. Yuan, J. J. Pestka, B. M. Hespenheide, L. A. Kuhn, J. E. Linz, and L. P. Hart (1999) "Identification of mimotope peptides which bind to the mycotoxin deoxynivalenol-specific monoclonal antibody" (pdf) Appl. Environ. Microbiol. 65, 3279-3286. D. Jacobs, L. A. Kuhn, and M. F. Thorpe (1999) "Flexible and Rigid Regions in Proteins" (pdf) in Rigidity Theory and Applications (M. F. Thorpe and P. M. Duxbury, eds.), Plenum Publishing, New York, pp. 357-384. V. Schnecke and L. A. Kuhn (1999) "Flexibly Screening for Molecules Interacting with Proteins" (pdf) in Rigidity Theory and Applications (M. F. Thorpe and P. M. Duxbury, eds.), Plenum Publishing, New York, pp. 385-400. V. Schnecke, C. A. Swanson, E. D. Getzoff, J. A. Tainer, and L. A. Kuhn (1998) "Screening a Peptidyl Database for Potential Ligands to Proteins with Side-chain Flexibility" (pdf) Proteins: Structure, Function, and Genetics, 33, 74-87. P. C. Sanschagrin and L. A. Kuhn (1998) "Cluster Analysis of Consensus Water Sites in Thrombin and Trypsin Shows Conservation Between Serine Proteases and Contributions to Ligand Specificity" (pdf) Protein Science, 7, 2054-2064. L. Craig, P. C. Sanschagrin, A. Rozek, S. Lackie, L. A. Kuhn, and J. K. Scott(1998) "The Role of Structure in Antibody Cross-Reactivity Between Peptides and Folded Proteins" (pdf) J. Mol. Biol., 281, 183-201. M. L. Raymer, W. F. Punch, E. D. Goodman, P. C. Sanschagrin, and L. A. Kuhn (1997) "Simultaneous Feature Scaling and Selection Using a Genetic Algorithm" (pdf) in Proceedings of the Seventh International Conference on Genetic Algorithms (T. Bäck, ed.), Morgan Kaufmann Publishers, San Francisco, pp. 561-567. M. L. Raymer, P.C. Sanschagrin, W. F. Punch, S. Venkataraman, E. D. Goodman, and L. A. Kuhn (1997) "Predicting Conserved Water-Mediated and Polar Ligand Interactions in Proteins Using a K-nearest-neighbors Genetic Algorithm" (pdf) J. Mol. Biol., 265, 445-464. M. L. Raymer, W. F. Punch, E. D. Goodman, and L. A. Kuhn (1996) "Genetic Programming for Improved Data Mining - Application to the Biochemistry of Protein Interactions" (pdf) in Genetic Programming 1996: Proceedings (J. R. Koza, D. E. Goldberg, D. B. Fogel, and R. L. Riolo, eds.), MIT Press, Cambridge, MA, pp. 275-381. L. A. Kuhn, C. A. Swanson, M. E. Pique, J. A. Tainer, and E. D. Getzoff (1995) "Atomic and Residue Hydrophilicity in the Context of Folded Protein Structures" (pdf) Proteins: Struct. Funct. Genet., 23, 536-547. L. A. Kuhn and C. L. M. J. Verlinde (1995) "Protein Structure-Based Drug Design" (pdf) Fall American Crystallographic Association Newsletter, pp. 23-24. L. Pond, L. A. Kuhn, M. R. Jackson, L. Teyton, M.-P. Schutze, J. A. Tainer, and P. A. Peterson (1995) "A Role for Acidic Residues in Di-leucine Motif-based Targeting to the Endocytic Pathway" (pdf) J. Biol. Chem. 270, 19989-19997.
